MonetDBLite for R
Today we are happy to present MonetDBLite for R, a fully embedded version of MonetDB that installs like any other R package. The database runs within the R process itself (like its namesake SQLite), greatly improving efficiency of data transfers. The package is available for Linux, Mac OS X and Windows (64 bit).
R and MonetDB fit together well, with statistical programming performed in R and data wrangling handled by MonetDB. For years, we have had the MonetDB.R R package, which allows to connect to a running MonetDB server, send queries and retrieve results. Due to the inefficiencies of shipping data around, MonetDB also supports User-Defined Functions implemented in R. However, both approaches require users to set up and maintain a separate MonetDB server. There have been numerous requests to simplify this process.
Installation of MonetDBLite is straightforward:
install.packages("MonetDBLite")
This will install MonetDBLite and the latest version of the MonetDB.R client package, which contains the DBI and dplyr backends. Here is a usage example with MonetDBLite:
library(DBI)
dbdir <- tempdir()
con <- dbConnect(MonetDBLite::MonetDBLite())
dbWriteTable(con, "mtcars", mtcars)
dbGetQuery(con, "SELECT MAX(mpg) FROM mtcars WHERE cyl = 8")
library(dplyr)
ms <- MonetDBLite::src_monetdblite(dbdir)
mt <- tbl(ms, "mtcars")
mt %>% filter(cyl == 8) %>% summarise(max(mpg))
Note that MonetDBLite will store the “mtcars” table on disk in the specified directory (a temporary one here). A later R session does not need to repeat writing the data to the database and can directly work with the stored persistent tables. This considerably reduces the startup time of analyses, especially as tables get large.
To showcase the power of MonetDBLite, we compared its performance against MonetDB with the socket-based R client and RSQLite. As a real-world benchmark, we use Anthony Damico’s scripts to analyse the “Home Mortgage Disclosure Act (HMDA)” dataset . These scripts generate SQL queries to accurately reproduce statistics found in Federal Financial Institutions Examination Council (FFIEC) publications. The used data from 2006 to 2011 contains 128 Million rows of data with 71 fields each. In CSV format, the full dataset occupies 56 GB of disk space. In addition, we have produced smaller versions of the dataset with 6M and 60M tuples by way of random sampling. We ran three experiments: CSV loading, querying and full table transfer into R.
All experiments report wall clock time and were run on a Linux desktop machine with 16GB of main memory and a 3.40GHz Intel i7-2600K CPU. The query and conversion timings are from “hot” runs. The following figure illustrate the outcome of the CSV loading experiment:
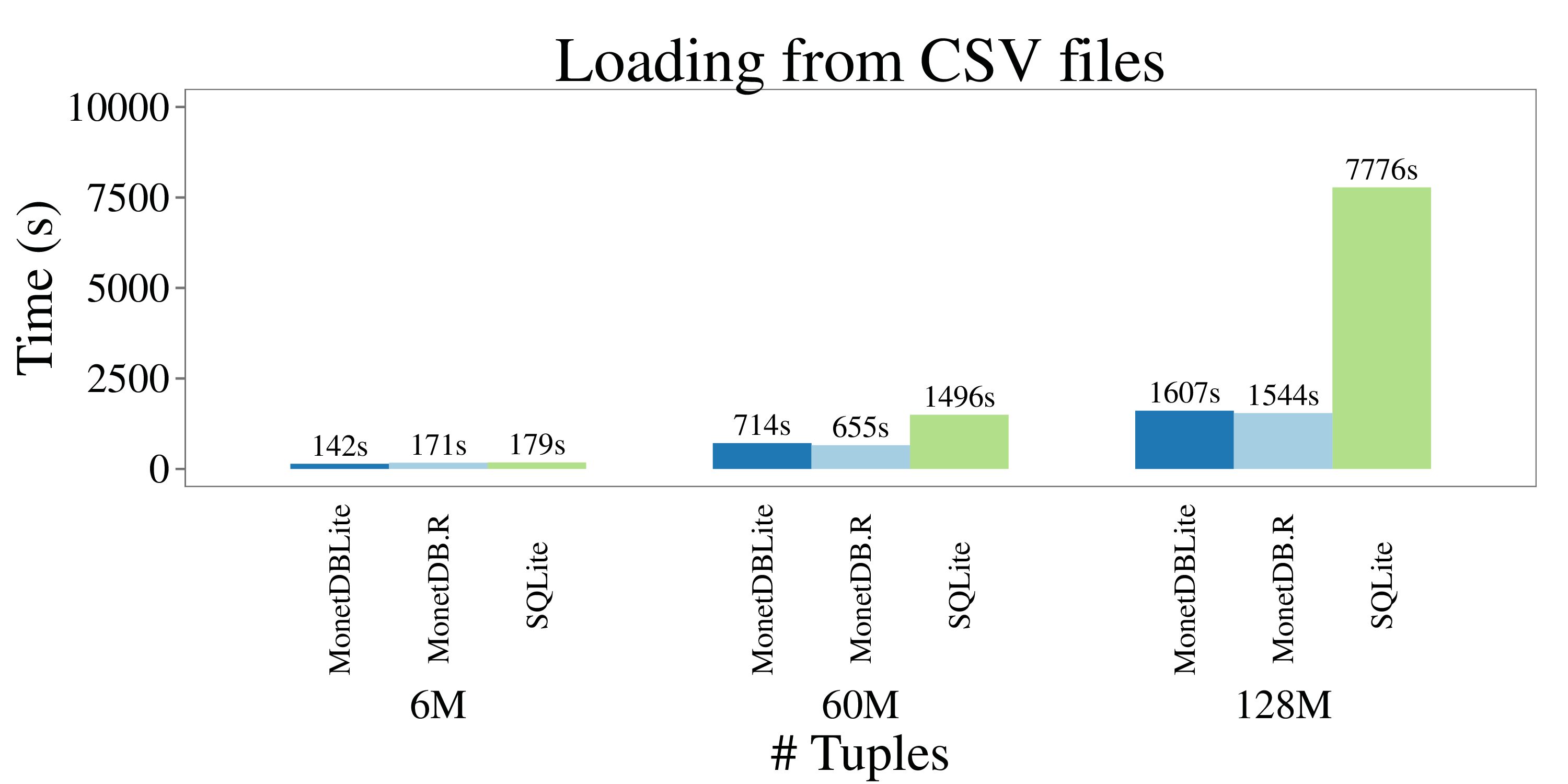
We can see how MonetDBLite and MonetDB.R are almost equal, which is of no surprise since they use the same CSV loader code. SQLite performs well except for the large dataset, where the operating system was forced to swap out virtual memory frequently.
Timings from running the HMDA queries are plotted in the following figure:
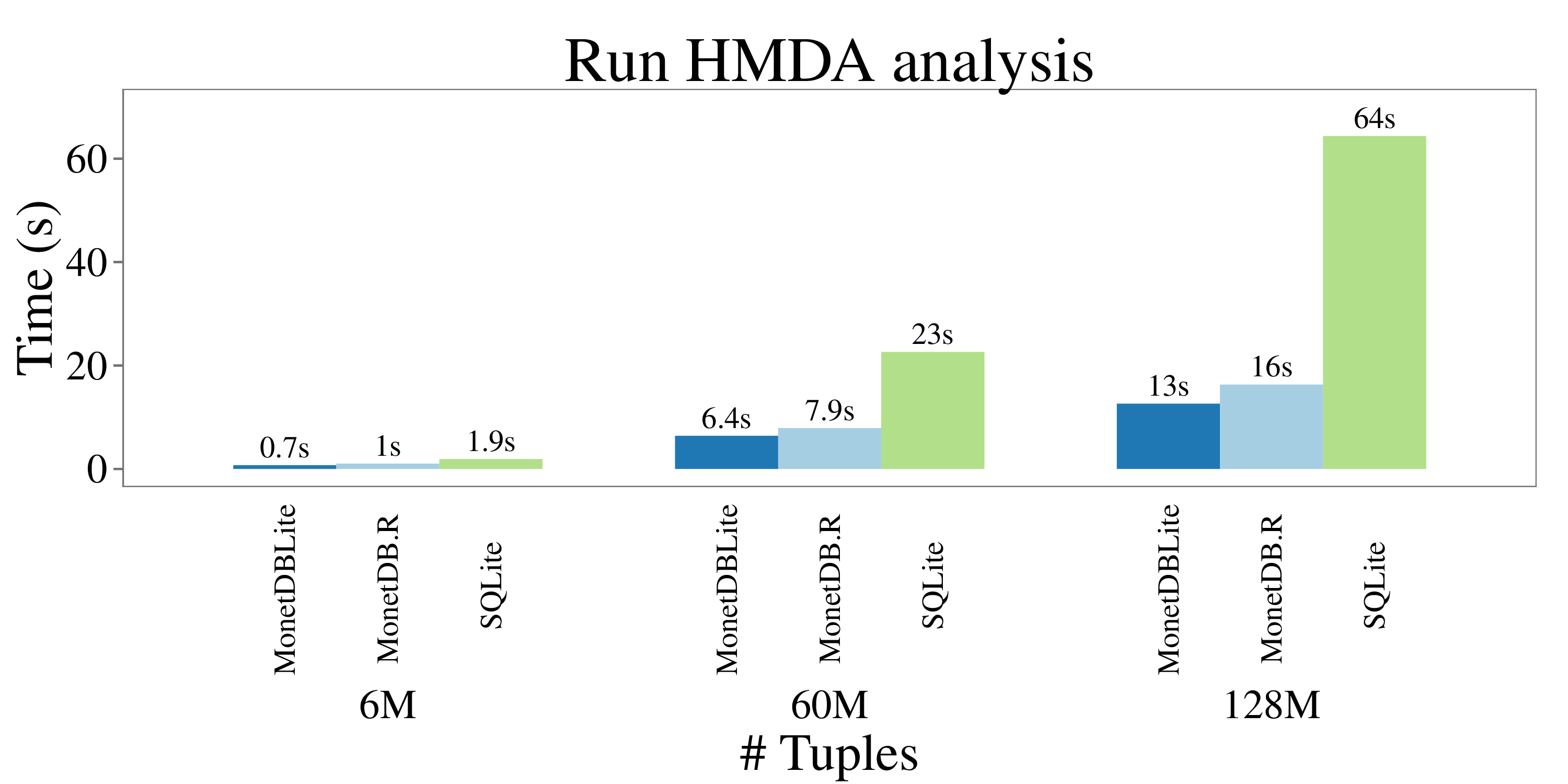
Since the query results only contain a few tuples, the socket overhead is small and the timings of MonetDBLite and MonetDB.R are again comparable. SQLite cannot benefit from the columnar storage model and performs significantly worse. **
Finally, another common use case is to retrieve an entire table from the database into a R data.frame. The next figure shows timings for reading the entire “hmda_11” table into R. **
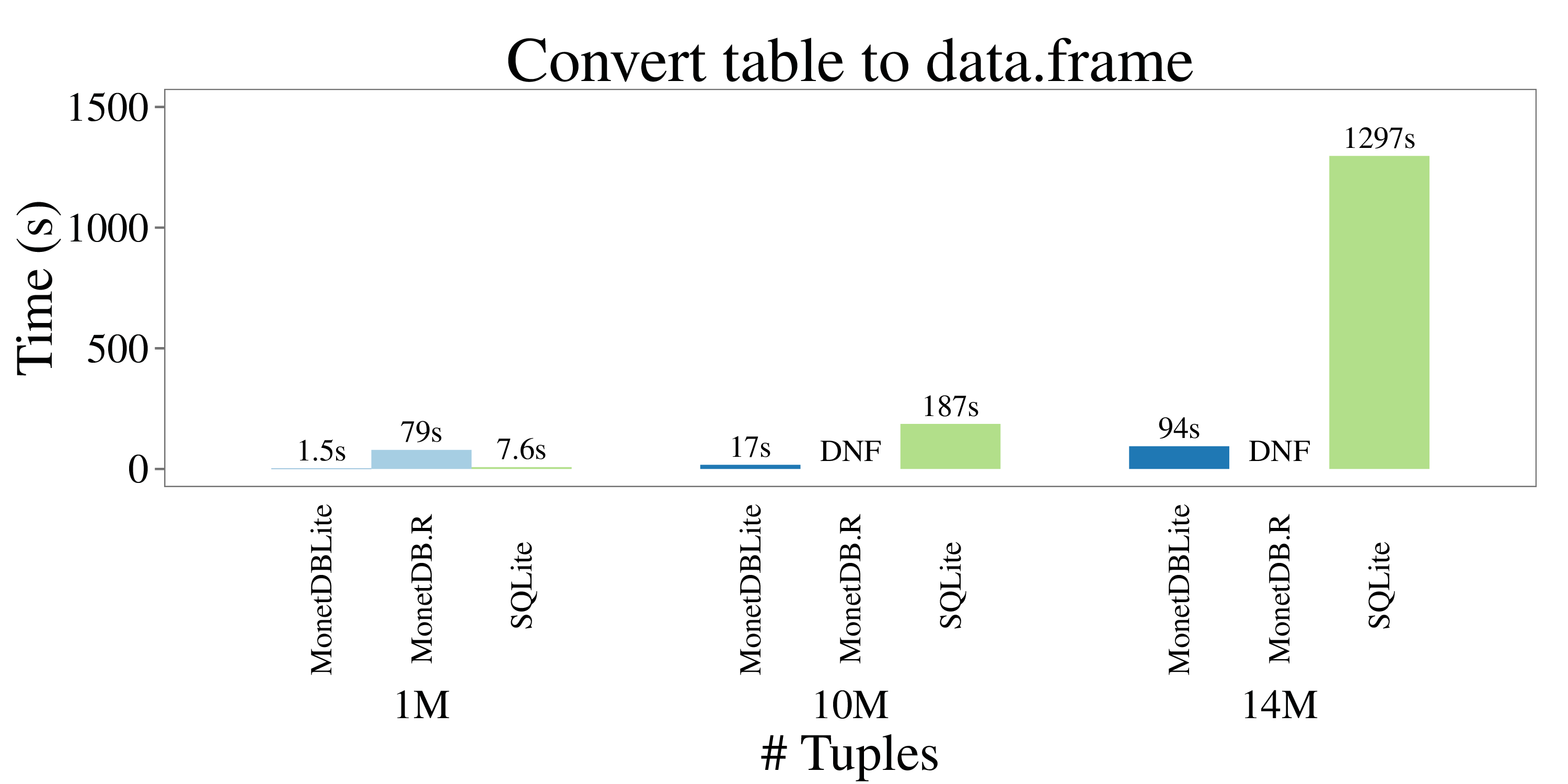
The socket-based MonetDB.R client is not able to perform well here. The client needs to convert the row-based wire protocol back to the colummnar format of data.frame objects and needs to perform extensive string parsing as well. Due to limitations in R, it did not finish (DNF) for the two larger data sets. MonetDBLite on the other hand shows stellar performance here, only taking 94 seconds to transfer 14M rows with 71 fields into R. This is largely due to their compatible columnar data representations. SQLite takes 13 times longer.
All scripts and data files used to produce these plots are available online..
We invite the interested public to try out MonetDBLite. If any issues arise, please report back to hannes@cwi.nl .
Update: MonetDBLite has been accepted to CRAN.
Update II: MonetDBLite is now on GitHub.